CTable Tutorial¶
CTable is a columnar compressed table built on top of blosc2.NDArray. It stores each column independently as a compressed array, giving you:
Compression — data lives compressed in RAM and on disk.
Schema — every column has a declared type and optional constraints.
Speed — bulk operations stay in NumPy; no row-by-row Python overhead.
Persistence — tables can be saved to and loaded from disk transparently.
This notebook walks through the full API, starting from the very basics and finishing with a real-world analysis of climate data across ten world cities.
[1]:
from dataclasses import dataclass
import matplotlib.pyplot as plt
import numpy as np
import blosc2
from blosc2 import CTable
Part 1 — The Basics¶
1.1 Defining a schema¶
Every CTable is typed. You define the schema with a plain Python @dataclass. Each field gets a spec — a blosc2 type that carries the NumPy dtype and optional constraints.
[2]:
@dataclass
class Sensor:
id: int = blosc2.field(blosc2.int32(ge=0))
location: str = blosc2.field(blosc2.string(max_length=16), default="")
temperature: float = blosc2.field(blosc2.float64(ge=-80, le=60), default=20.0)
active: bool = blosc2.field(blosc2.bool(), default=True)
# Create an empty in-memory table
t = CTable(Sensor, expected_size=50)
print(f"Empty table: {len(t)} rows, columns: {t.col_names}")
Empty table: 0 rows, columns: ['id', 'location', 'temperature', 'active']
1.2 Appending rows¶
append() adds one row at a time. The row is validated against the schema before writing.
[3]:
t.append(Sensor(id=0, location="roof", temperature=22.5, active=True))
t.append(Sensor(id=1, location="basement", temperature=18.1, active=True))
t.append(Sensor(id=2, location="outdoor", temperature=-3.2, active=False))
print(t)
id location temperature active
int32 U16 (Unicode, max 16 chars) float64 bool
──────────── ───────────────────────────── ───────────────── ────────
0 roof 22.5 True
1 basement 18.1 True
2 outdoor -3.2 False
──────────── ───────────────────────────── ───────────────── ────────
3 rows × 4 columns
Constraints are enforced — trying to insert a temperature above 60 °C raises an error:
[4]:
try:
t.append(Sensor(id=99, location="sun", temperature=9999.0, active=True))
except Exception as e:
print(f"Validation error: {e}")
Validation error: 1 validation error for _Validator_Sensor
temperature
Input should be less than or equal to 60 [type=less_than_equal, input_value=9999.0, input_type=float]
For further information visit https://errors.pydantic.dev/2.13/v/less_than_equal
1.3 Bulk loading with extend()¶
extend() accepts a list of tuples or a structured NumPy array. It is much faster than calling append() in a loop.
[5]:
bulk = [
(3, "lab-A", 20.0, True),
(4, "lab-B", 21.5, True),
(5, "server", 35.8, True),
(6, "garden", -1.0, False),
]
t.extend(bulk)
print(f"After extend: {len(t)} rows")
print(t)
After extend: 7 rows
id location temperature active
int32 U16 (Unicode, max 16 chars) float64 bool
──────────── ───────────────────────────── ───────────────── ────────
0 roof 22.5 True
1 basement 18.1 True
2 outdoor -3.2 False
3 lab-A 20.0 True
4 lab-B 21.5 True
5 server 35.8 True
6 garden -1.0 False
──────────── ───────────────────────────── ───────────────── ────────
7 rows × 4 columns
1.5 Columns as first-class objects¶
Access a column with table["name"] or table.name. Columns are lazy — they only decompress data when you ask for it.
[7]:
temps = t["temperature"]
print(f"dtype : {temps.dtype}")
print(f"min : {temps.min():.1f} °C")
print(f"max : {temps.max():.1f} °C")
print(f"mean : {temps.mean():.1f} °C")
print(f"as numpy: {temps[:]}")
dtype : float64
min : -3.2 °C
max : 35.8 °C
mean : 16.2 °C
as numpy: [22.5 18.1 -3.2 20. 21.5 35.8 -1. ]
1.6 Computed columns¶
A CTable can also expose computed columns: read-only columns backed by a lazy expression over stored columns. They use no extra storage, update automatically after appends/deletes, and participate in display, filtering, sorting, and aggregates.
[8]:
t.add_computed_column("temperature_f", "temperature * 9 / 5 + 32")
print(t.select(["location", "temperature", "temperature_f"]))
print(f"\nMean temperature in °F: {t['temperature_f'].mean():.1f} °F")
# Use the computed column in a query
warm_f = t.where(t.temperature_f > 70)
print("\nRows above 70 °F:")
print(warm_f.select(["location", "temperature", "temperature_f"]))
location temperature temperature_f
U16 (Unicode, max 16 chars) float64 float64
───────────────────────────── ───────────────── ─────────────────
roof 22.5 72.5
basement 18.1 64.58
outdoor -3.2 26.24000000000…
lab-A 20.0 68.0
lab-B 21.5 70.7
server 35.8 96.44
garden -1.0 30.2
───────────────────────────── ───────────────── ─────────────────
7 rows × 3 columns
Mean temperature in °F: 61.2 °F
Rows above 70 °F:
location temperature temperature_f
U16 (Unicode, max 16 chars) float64 float64
───────────────────────────── ───────────────── ─────────────────
roof 22.5 72.5
lab-B 21.5 70.7
server 35.8 96.44
───────────────────────────── ───────────────── ─────────────────
3 rows × 3 columns
Part 2 — The Climate Dataset¶
Enough warm-up. Let’s do something real.
We will simulate one full year of daily weather readings for 10 world cities. Each row is one day at one city: temperature, humidity, wind speed, atmospheric pressure.
City |
Climate |
Twist |
|---|---|---|
Madrid |
Mediterranean |
Scorching summers, mild winters |
London |
Temperate oceanic |
Famously grey and damp |
Beijing |
Continental |
Brutal winters, hot summers |
New York |
Humid continental |
Four very distinct seasons |
Tokyo |
Humid subtropical |
Warm and very humid summers |
Sydney |
Oceanic (S. hemisphere) |
Seasons are flipped! |
Cairo |
Hot desert |
Basically always hot |
Moscow |
Subarctic |
Coldest city in the dataset |
Mumbai |
Tropical |
Hot and humid all year |
São Paulo |
Tropical highland |
Warm, rainy, south hemisphere |
[9]:
@dataclass
class WeatherReading:
city: str = blosc2.field(blosc2.string(max_length=16))
day: int = blosc2.field(blosc2.int16(ge=1, le=365), default=1)
temperature: float = blosc2.field(blosc2.float32(ge=-80.0, le=60.0), default=20.0)
humidity: float = blosc2.field(blosc2.float32(ge=0.0, le=100.0), default=50.0)
wind_speed: float = blosc2.field(blosc2.float32(ge=0.0, le=200.0), default=0.0)
pressure: float = blosc2.field(blosc2.float32(ge=800.0, le=1100.0), default=1013.0)
[10]:
# Climate profile for each city:
# mean_temp : annual mean temperature (°C)
# amplitude : half the annual temperature swing (°C)
# peak_day : day of year with the highest temperature
# (196 ≈ July 15 for N. hemisphere, 15 ≈ Jan 15 for S. hemisphere)
# humidity : annual mean relative humidity (%)
# wind : mean wind speed (km/h)
# pressure : mean atmospheric pressure (hPa)
CITY_PROFILES = {
"Madrid": {
"mean_temp": 15.0,
"amplitude": 13.0,
"peak_day": 196,
"humidity": 45,
"wind": 12,
"pressure": 1010,
},
"London": {
"mean_temp": 11.0,
"amplitude": 7.0,
"peak_day": 196,
"humidity": 75,
"wind": 15,
"pressure": 1013,
},
"Beijing": {
"mean_temp": 12.0,
"amplitude": 16.0,
"peak_day": 196,
"humidity": 55,
"wind": 10,
"pressure": 1012,
},
"New York": {
"mean_temp": 13.0,
"amplitude": 14.0,
"peak_day": 196,
"humidity": 65,
"wind": 14,
"pressure": 1013,
},
"Tokyo": {
"mean_temp": 15.0,
"amplitude": 12.0,
"peak_day": 196,
"humidity": 72,
"wind": 11,
"pressure": 1014,
},
"Sydney": {
"mean_temp": 18.0,
"amplitude": 8.0,
"peak_day": 15,
"humidity": 65,
"wind": 16,
"pressure": 1012,
},
"Cairo": {
"mean_temp": 22.0,
"amplitude": 14.0,
"peak_day": 196,
"humidity": 35,
"wind": 8,
"pressure": 1014,
},
"Moscow": {
"mean_temp": 5.0,
"amplitude": 18.0,
"peak_day": 196,
"humidity": 70,
"wind": 10,
"pressure": 1015,
},
"Mumbai": {
"mean_temp": 28.0,
"amplitude": 4.0,
"peak_day": 196,
"humidity": 80,
"wind": 12,
"pressure": 1011,
},
"Sao Paulo": {
"mean_temp": 22.0,
"amplitude": 5.0,
"peak_day": 15,
"humidity": 75,
"wind": 8,
"pressure": 1016,
},
}
rng = np.random.default_rng(42)
days = np.arange(1, 366, dtype=np.int16)
all_rows = []
for city, p in CITY_PROFILES.items():
seasonal = p["amplitude"] * np.cos(2 * np.pi * (days - p["peak_day"]) / 365)
temps = (p["mean_temp"] + seasonal + rng.normal(0, 2.0, 365)).clip(-80, 60).astype(np.float32)
humidity = (p["humidity"] + rng.normal(0, 8.0, 365)).clip(0, 100).astype(np.float32)
wind = (p["wind"] + rng.exponential(4.0, 365)).clip(0, 200).astype(np.float32)
pressure = (p["pressure"] + rng.normal(0, 5.0, 365)).clip(800, 1100).astype(np.float32)
for i, d in enumerate(days):
all_rows.append(
(city, int(d), float(temps[i]), float(humidity[i]), float(wind[i]), float(pressure[i]))
)
climate = CTable(WeatherReading, new_data=all_rows, validate=False, expected_size=len(all_rows))
print(f"Climate table: {len(climate):,} rows × {climate.ncols} columns")
print(f"Compressed: {climate.cbytes / 1024:.1f} KB (uncompressed: {climate.nbytes / 1024:.1f} KB)")
print(climate)
Climate table: 3,650 rows × 6 columns
Compressed: 42.1 KB (uncompressed: 295.8 KB)
city day temperature humidity wind_speed pressure
U16 (Unicode, max 16 chars) int16 float32 float32 float32 float32
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
Madrid 1 2.909233808… 56.85164260… 14.76450634… 1014.201110…
Madrid 2 0.173932701… 39.05129241… 12.57167434… 1017.437011…
Madrid 3 3.712681531… 38.42200088… 12.35102844… 1008.641601…
Madrid 4 4.054576396… 46.61845016… 14.67175292… 1004.238769…
Madrid 5 -1.76315498… 51.75508117… 18.27811622… 1008.797790…
Madrid 6 -0.49616417… 45.09140777… 12.12416076… 1010.510131…
Madrid 7 2.336975336… 55.63168334… 12.29123497… 1010.394958…
Madrid 8 1.425737023… 51.85435104… 12.44658184… 1015.668395…
Madrid 9 2.005382537… 51.73456192… 12.67978572… 1008.193664…
Madrid 10 0.317500203… 49.43293380… 15.99008655… 1011.760192…
... ... ... ... ... ...
Sao Paulo 356 24.10465049… 79.09764862… 11.10457992… 1014.604797…
Sao Paulo 357 28.06665992… 75.45437622… 8.651307106… 1021.078308…
Sao Paulo 358 25.48140907… 70.06164550… 9.325889587… 1010.540771…
Sao Paulo 359 25.73676872… 85.70543670… 8.712495803… 1011.278442…
Sao Paulo 360 25.67271423… 76.63825988… 10.50052738… 1019.007934…
Sao Paulo 361 26.86038208… 80.04640960… 11.57989692… 1013.804260…
Sao Paulo 362 24.26036834… 86.33271026… 11.86164474… 1008.734680…
Sao Paulo 363 22.30093574… 86.20143127… 13.39677143… 1022.754089…
Sao Paulo 364 27.72695350… 79.63828277… 17.80615234… 1010.502014…
Sao Paulo 365 22.60712432… 72.57873535… 9.853503227… 1015.002868…
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
3,650 rows × 6 columns (3,630 rows hidden)
Part 3 — Querying¶
3.1 Filtering rows with where()¶
where() takes a boolean expression built from column comparisons and returns a view — a lightweight object that shares the underlying data without copying it.
[11]:
# All days where temperature exceeded 35 °C (any city)
very_hot = climate.where(climate.temperature > 35)
print(f"Days above 35 °C: {len(very_hot)} ({len(very_hot) / len(climate) * 100:.1f}% of all readings)")
print(very_hot.head(8))
Days above 35 °C: 49 (1.3% of all readings)
city day temperature humidity wind_speed pressure
U16 (Unicode, max 16 chars) int16 float32 float32 float32 float32
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
Cairo 154 35.72507095… 39.59734344… 10.80750942… 1010.431884…
Cairo 157 35.66441726… 38.08246231… 9.141173362… 1016.997558…
Cairo 158 35.80884170… 34.47190475… 12.61370849… 1016.754882…
Cairo 162 35.91463088… 33.77049636… 20.62659454… 1008.747253…
Cairo 163 36.98370361… 31.69925498… 15.52884197… 1010.481750…
Cairo 165 37.55741119… 35.59812164… 9.190578460… 1014.468322…
Cairo 169 36.81953430… 40.87221145… 15.42489051… 1024.706420…
Cairo 170 37.21762847… 36.48490905… 12.23543548… 1012.218139…
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
8 rows × 6 columns
[12]:
# Moscow in winter (below freezing)
moscow_frozen = climate.where((climate.city == "Moscow") & (climate.temperature < 0))
print(f"Moscow below freezing: {len(moscow_frozen)} days out of 365")
print(moscow_frozen.head())
Moscow below freezing: 148 days out of 365
city day temperature humidity wind_speed pressure
U16 (Unicode, max 16 chars) int16 float32 float32 float32 float32
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
Moscow 1 -13.5099849… 76.91526794… 10.18380069… 1006.785095…
Moscow 2 -13.0531520… 77.78500366… 20.35687637… 1010.101074…
Moscow 3 -12.9442214… 81.18754577… 16.77510261… 1020.993286…
Moscow 4 -12.8625192… 73.40472412… 13.44744586… 1013.957031…
Moscow 5 -10.4717388… 69.11986541… 10.80644416… 1016.391967…
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
5 rows × 6 columns
3.2 Column projection with select()¶
select() returns a view with only the columns you need — no data is copied.
[13]:
# Just city, day, and temperature — useful before exporting or computing stats
slim = climate.select(["city", "day", "temperature"])
print(slim.head(6))
city day temperature
U16 (Unicode, max 16 chars) int16 float32
───────────────────────────── ────────── ──────────────
Madrid 1 2.909233808…
Madrid 2 0.173932701…
Madrid 3 3.712681531…
Madrid 4 4.054576396…
Madrid 5 -1.76315498…
Madrid 6 -0.49616417…
───────────────────────────── ────────── ──────────────
6 rows × 3 columns
3.3 Sorting¶
sort_by() returns a sorted copy by default (or sorts in-place with inplace=True). Multi-column sorting is supported — primary key first.
[14]:
# Which were the 10 hottest days across all cities?
hottest = climate.sort_by("temperature", ascending=False)
print("Top 10 hottest days (any city):")
print(hottest.head(10))
Top 10 hottest days (any city):
city day temperature humidity wind_speed pressure
U16 (Unicode, max 16 chars) int16 float32 float32 float32 float32
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
Cairo 225 39.74710083… 40.93272018… 11.85465049… 1011.177551…
Cairo 184 39.47347640… 36.25406646… 8.386101722… 1009.952026…
Cairo 195 39.28937149… 26.06153678… 9.814800262… 1025.788452…
Cairo 205 38.39984130… 47.71873855… 21.52345466… 1014.675598…
Cairo 213 38.25638198… 38.09883117… 17.99407577… 1013.649597…
Cairo 218 37.88246536… 29.69907951… 11.12999248… 1015.036682…
Cairo 185 37.75819396… 30.60622787… 9.116196632… 1015.961669…
Cairo 165 37.55741119… 35.59812164… 9.190578460… 1014.468322…
Cairo 177 37.33380889… 23.67673683… 8.815520286… 1013.045227…
Cairo 170 37.21762847… 36.48490905… 12.23543548… 1012.218139…
───────────────────────────── ────────── ────────────── ────────────── ────────────── ──────────────
10 rows × 6 columns
[15]:
# Multi-column sort: primary key = city (A→Z), secondary = temperature (hottest first)
# This lets you see each city's hottest day at a glance
by_city_temp = climate.sort_by(["city", "temperature"], ascending=[True, False])
print("Sorted by city (asc) then temperature (desc):")
print(by_city_temp.select(["city", "day", "temperature", "humidity"]).head(20))
Sorted by city (asc) then temperature (desc):
city day temperature humidity
U16 (Unicode, max 16 chars) int16 float32 float32
───────────────────────────── ────────── ────────────── ──────────────
Beijing 207 33.77753448… 53.55015182…
Beijing 222 31.76755332… 55.32269287…
Beijing 205 31.75275421… 56.29219818…
Beijing 214 30.79148101… 63.60578536…
Beijing 178 30.47000694… 53.21742248…
Beijing 202 30.44628334… 65.57708740…
Beijing 203 30.01535034… 60.08674621…
Beijing 164 29.65973281… 43.95068740…
Beijing 177 29.58276939… 37.89999389…
Beijing 196 29.33683586… 54.66562271…
Beijing 188 29.22382354… 57.12422943…
Beijing 212 29.06817436… 43.84931945…
Beijing 191 29.05400466… 45.30105209…
Beijing 189 28.97620391… 55.71607208…
Beijing 181 28.96719360… 51.00849914…
Beijing 204 28.92338752… 45.47613143…
Beijing 179 28.87882232… 64.10365295…
Beijing 193 28.80445098… 52.74388504…
Beijing 195 28.73704147… 66.52650451…
Beijing 160 28.67715644… 61.17769241…
───────────────────────────── ────────── ────────────── ──────────────
20 rows × 4 columns
Part 4 — Aggregates and Statistics¶
4.1 Per-city mean temperature¶
[16]:
print(f"{'City':<12} {'Mean temp':>10} {'Min':>7} {'Max':>7} {'Std':>7}")
print("-" * 50)
for city in CITY_PROFILES:
v = climate.where(climate.city == city)
col = v["temperature"]
print(f"{city:<12} {col.mean():>9.1f}° {col.min():>6.1f}° {col.max():>6.1f}° {col.std():>6.1f}°")
City Mean temp Min Max Std
--------------------------------------------------
Madrid 15.0° -1.8° 31.4° 9.3°
London 10.8° -0.3° 22.7° 5.3°
Beijing 12.1° -9.1° 33.8° 11.5°
New York 13.0° -4.4° 30.9° 10.2°
Tokyo 15.1° -0.2° 31.0° 8.5°
Sydney 17.8° 4.7° 30.9° 5.9°
Cairo 21.9° 2.8° 39.7° 10.1°
Moscow 5.0° -17.5° 26.3° 12.9°
Mumbai 27.9° 18.4° 36.6° 3.5°
Sao Paulo 21.9° 12.7° 30.9° 4.1°
4.2 describe() — full summary in one call¶
[17]:
# describe() on a select() view — only numeric columns
climate.select(["temperature", "humidity", "wind_speed", "pressure"]).describe()
CTable 3,650 rows × 4 cols
temperature [float32]
count : 3,650
mean : 16.04
std : 10.72
min : -17.54
max : 39.75
humidity [float32]
count : 3,650
mean : 63.48
std : 16.02
min : 8.894
max : 99.81
wind_speed [float32]
count : 3,650
mean : 15.63
std : 4.874
min : 8.005
max : 47.48
pressure [float32]
count : 3,650
mean : 1013
std : 5.328
min : 991.1
max : 1036
4.3 Covariance matrix¶
cov() requires all columns to be numeric (int, float, or bool). It returns a standard numpy.ndarray.
[18]:
numeric = climate.select(["temperature", "humidity", "wind_speed", "pressure"])
cov = numeric.cov()
labels = ["temp", "humidity", "wind", "pressure"]
col_w = 12
print("Covariance matrix (all cities, full year):")
print(" " * 10 + "".join(f"{lbl:>{col_w}}" for lbl in labels))
for i, lbl in enumerate(labels):
print(f"{lbl:<10}" + "".join(f"{cov[i, j]:>{col_w}.3f}" for j in range(4)))
# And the correlation matrix for easier interpretation
corr = np.corrcoef(np.stack([numeric[c][:] for c in ["temperature", "humidity", "wind_speed", "pressure"]]))
print("\nCorrelation matrix:")
print(" " * 10 + "".join(f"{lbl:>{col_w}}" for lbl in labels))
for i, lbl in enumerate(labels):
print(f"{lbl:<10}" + "".join(f"{corr[i, j]:>{col_w}.3f}" for j in range(4)))
Covariance matrix (all cities, full year):
temp humidity wind pressure
temp 114.963 0.018 -3.523 -0.207
humidity 0.018 256.861 10.773 6.652
wind -3.523 10.773 23.760 -2.650
pressure -0.207 6.652 -2.650 28.394
Correlation matrix:
temp humidity wind pressure
temp 1.000 0.000 -0.067 -0.004
humidity 0.000 1.000 0.138 0.078
wind -0.067 0.138 1.000 -0.102
pressure -0.004 0.078 -0.102 1.000
Part 5 — Analysis: Summer in Madrid¶
Summer in the northern hemisphere runs roughly from the summer solstice (day 172, June 21) to the autumnal equinox (day 264, September 22).
Let’s zoom in on Madrid during those months and compare it with a few other cities.
[19]:
SUMMER_START = 172 # June 21
SUMMER_END = 264 # September 22
madrid = climate.where(climate.city == "Madrid")
madrid_summer = madrid.where((madrid.day >= SUMMER_START) & (madrid.day <= SUMMER_END))
print(f"Madrid summer readings : {len(madrid_summer)} days")
print(f" mean temperature : {madrid_summer.temperature.mean():.1f} °C")
print(f" max temperature : {madrid_summer.temperature.max():.1f} °C")
print(f" mean humidity : {madrid_summer['humidity'].mean():.1f} %")
print(f" mean wind speed : {madrid_summer['wind_speed'].mean():.1f} km/h")
Madrid summer readings : 93 days
mean temperature : 25.8 °C
max temperature : 31.4 °C
mean humidity : 43.8 %
mean wind speed : 15.8 km/h
[20]:
# Compare summer stats across several cities
compare_cities = ["Madrid", "London", "Cairo", "Moscow", "Tokyo", "Sydney"]
print(f"{'City':<12} {'Summer mean':>12} {'Summer max':>11} {'Summer humidity':>16}")
print("-" * 58)
for city in compare_cities:
v = climate.where(climate.city == city)
# For Sydney (S. hemisphere) 'summer' is Jan-Mar, i.e. days 1-80 or 355-365
if city == "Sydney":
s = v.where((v.day <= 80) | (v.day >= 355))
label = "(S. summer)"
else:
s = v.where((v.day >= SUMMER_START) & (v.day <= SUMMER_END))
label = ""
mean_t = s["temperature"].mean()
max_t = s["temperature"].max()
mean_h = s["humidity"].mean()
print(f"{city:<12} {mean_t:>10.1f}°C {max_t:>9.1f}°C {mean_h:>14.1f}% {label}")
City Summer mean Summer max Summer humidity
----------------------------------------------------------
Madrid 25.8°C 31.4°C 43.8%
London 16.5°C 22.7°C 74.6%
Cairo 33.5°C 39.7°C 34.4%
Moscow 20.1°C 26.3°C 69.3%
Tokyo 25.1°C 31.0°C 73.0%
Sydney 24.6°C 30.9°C 63.8% (S. summer)
[21]:
# Top 10 hottest days in Madrid across the whole year
# Sort the full table, then filter — views cannot be sorted directly
hottest_all = climate.sort_by("temperature", ascending=False)
madrid_sorted = hottest_all.where(hottest_all.city == "Madrid")
print("10 hottest days in Madrid:")
print(madrid_sorted.select(["city", "day", "temperature", "humidity"]).head(10))
10 hottest days in Madrid:
city day temperature humidity
U16 (Unicode, max 16 chars) int16 float32 float32
───────────────────────────── ────────── ────────────── ──────────────
Madrid 191 31.39920806… 42.54333496…
Madrid 190 31.23257637… 44.30324554…
Madrid 227 31.22744178… 46.99229049…
Madrid 194 30.91518402… 35.04422760…
Madrid 186 30.87937355… 48.08030319…
Madrid 202 30.74568367… 43.72281265…
Madrid 177 30.46902275… 38.39016342…
Madrid 163 30.21517944… 46.05188751…
Madrid 181 30.18102455… 43.72652053…
Madrid 184 29.93619918… 50.65479660…
───────────────────────────── ────────── ────────────── ──────────────
10 rows × 4 columns
5.1 Plotting: temperature over the year¶
Let’s visualise the full annual temperature cycle for a few contrasting cities.
[22]:
plot_cities = {
"Madrid": "#e63946",
"London": "#457b9d",
"Moscow": "#2d6a4f",
"Cairo": "#f4a261",
"Sydney": "#a8dadc",
}
fig, ax = plt.subplots(figsize=(12, 5))
for city, color in plot_cities.items():
v = climate.where(climate.city == city)
d = v.day[:].astype(int)
t = v["temperature"][:]
order = np.argsort(d)
ax.plot(d[order], t[order], label=city, color=color, linewidth=1.5, alpha=0.85)
ax.axvspan(SUMMER_START, SUMMER_END, alpha=0.10, color="gold", label="N. summer")
ax.set_xlabel("Day of year")
ax.set_ylabel("Temperature (°C)")
ax.set_title("Daily temperature — selected cities")
ax.legend(loc="upper left")
ax.grid(True, linestyle="--", alpha=0.4)
plt.tight_layout()
plt.show()
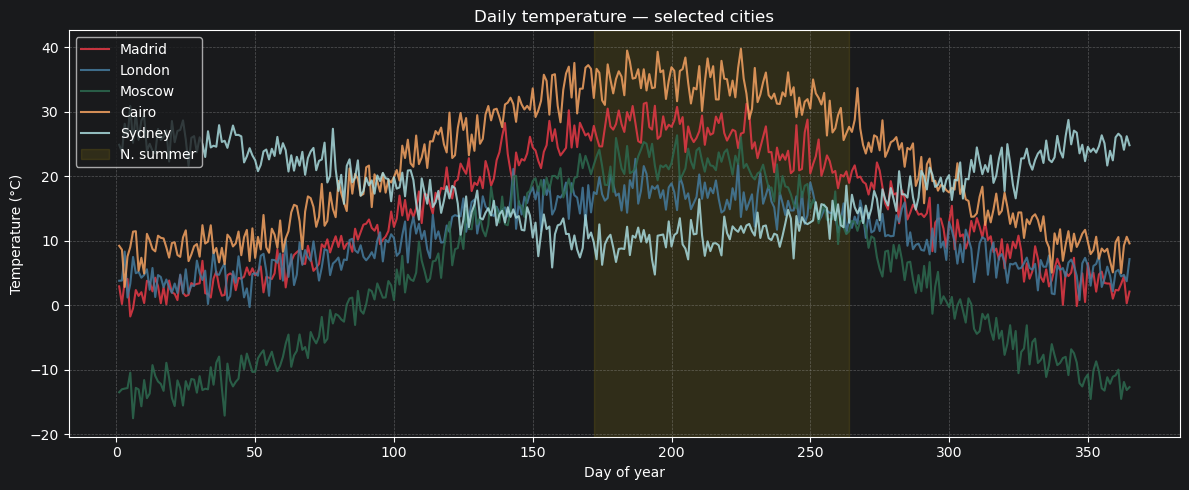
5.2 Summer temperature distribution — Madrid vs London¶
A simple histogram comparison of how often each city exceeds different temperature thresholds.
[23]:
madrid_s = climate.where(
(climate.city == "Madrid") & (climate.day >= SUMMER_START) & (climate.day <= SUMMER_END)
)["temperature"][:]
london_s = climate.where(
(climate.city == "London") & (climate.day >= SUMMER_START) & (climate.day <= SUMMER_END)
)["temperature"][:]
fig, ax = plt.subplots(figsize=(9, 4))
bins = np.linspace(0, 45, 30)
ax.hist(madrid_s, bins=bins, alpha=0.7, color="#e63946", label="Madrid")
ax.hist(london_s, bins=bins, alpha=0.7, color="#457b9d", label="London")
ax.axvline(
madrid_s.mean(),
color="#e63946",
linestyle="--",
linewidth=1.5,
label=f"Madrid mean {madrid_s.mean():.1f}°C",
)
ax.axvline(
london_s.mean(),
color="#457b9d",
linestyle="--",
linewidth=1.5,
label=f"London mean {london_s.mean():.1f}°C",
)
ax.set_xlabel("Temperature (°C)")
ax.set_ylabel("Days")
ax.set_title("Summer temperature distribution — Madrid vs London")
ax.legend()
ax.grid(True, linestyle="--", alpha=0.4)
plt.tight_layout()
plt.show()
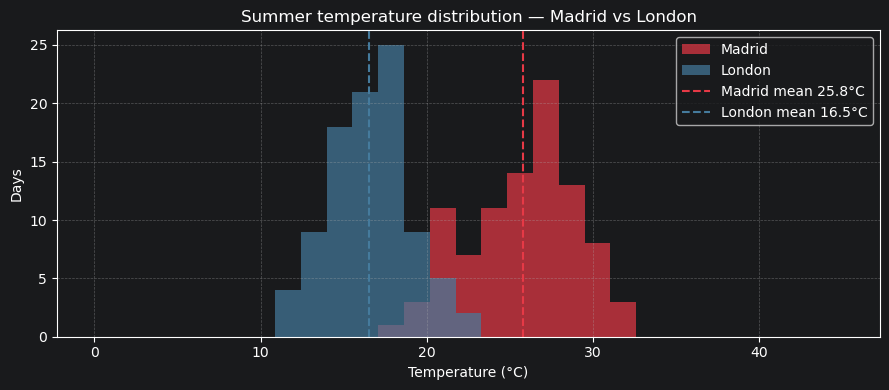
5.3 Mean summer temperature — all cities ranked¶
[24]:
city_summer_means = {}
for city in CITY_PROFILES:
v = climate.where(climate.city == city)
if city == "Sydney" or city == "Sao Paulo":
s = v.where((v.day <= 80) | (v.day >= 355))
else:
s = v.where((v.day >= SUMMER_START) & (v.day <= SUMMER_END))
city_summer_means[city] = s["temperature"].mean()
sorted_cities = sorted(city_summer_means.items(), key=lambda x: x[1], reverse=True)
names = [c for c, _ in sorted_cities]
means = [m for _, m in sorted_cities]
colors = ["#e63946" if m > 30 else "#f4a261" if m > 20 else "#457b9d" for m in means]
fig, ax = plt.subplots(figsize=(10, 4))
bars = ax.barh(names, means, color=colors, edgecolor="white")
ax.bar_label(bars, fmt="%.1f °C", padding=4)
ax.set_xlabel("Mean summer temperature (°C)")
ax.set_title("Cities ranked by summer heat")
ax.set_xlim(0, max(means) * 1.15)
ax.grid(True, axis="x", linestyle="--", alpha=0.4)
plt.tight_layout()
plt.show()
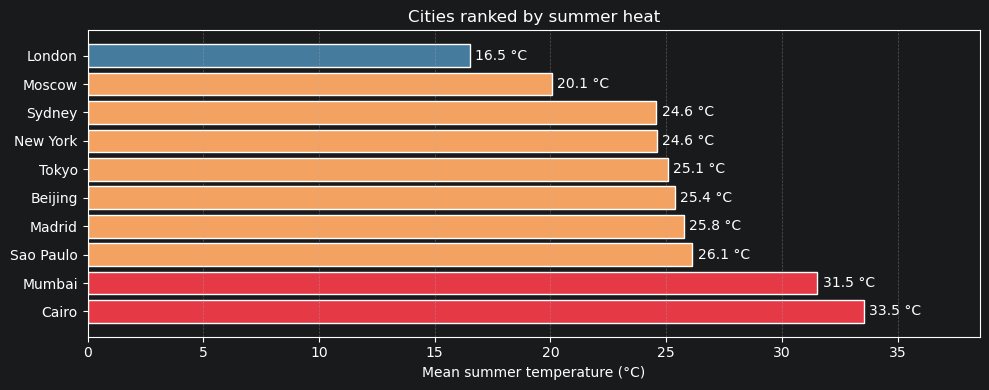
Part 6 — Mutations¶
CTable supports structural and value mutations: adding/dropping columns, deleting rows, sorting in place.
[25]:
# Add a 'feels_like' column: temperature adjusted for wind chill (simplified)
climate.add_column("feels_like", blosc2.float32(), default=0.0)
temp = climate.temperature[:]
wind = climate["wind_speed"][:]
# Simple wind-chill approximation (only meaningful below 10°C)
feels = np.where(temp < 10, temp - wind * 0.15, temp).astype(np.float32)
climate["feels_like"].assign(feels)
print("Table with feels_like column:")
print(climate.head(5))
print(f"\nColdest 'feels like' day: {climate['feels_like'].min():.1f} °C")
Table with feels_like column:
city day temperature humidity wind_speed pressure feels_like
U16 (Unicode, max 16 chars) int16 float32 float32 float32 float32 float32
───────────────────────────── ────────── ────────────── ────────────── ────────────── ────────────── ──────────────
Madrid 1 2.909233808… 56.85164260… 14.76450634… 1014.201110… 0.694557666…
Madrid 2 0.173932701… 39.05129241… 12.57167434… 1017.437011… -1.71181857…
Madrid 3 3.712681531… 38.42200088… 12.35102844… 1008.641601… 1.860027194…
Madrid 4 4.054576396… 46.61845016… 14.67175292… 1004.238769… 1.853813409…
Madrid 5 -1.76315498… 51.75508117… 18.27811622… 1008.797790… -4.50487232…
───────────────────────────── ────────── ────────────── ────────────── ────────────── ────────────── ──────────────
5 rows × 7 columns
Coldest 'feels like' day: -19.2 °C
Part 7 — Persistence¶
CTable can live on disk in two Blosc2 container formats:
.b2d: a directory-backed store. This is the best default for local read/write workflows..b2z: a single zip-backed store. This is compact and convenient for sharing or archiving, and is typically opened read-only.
Both formats keep the table columns compressed. Use open() for persistent/on-disk access and load() when you want an in-memory copy.
[26]:
import os
import shutil
import tempfile
tmpdir = tempfile.mkdtemp(prefix="blosc2_climate_")
disk_path = f"{tmpdir}/climate.b2d"
zip_path = f"{tmpdir}/climate.b2z"
unpacked_path = f"{tmpdir}/climate-unpacked.b2d"
compact_zip_path = f"{tmpdir}/climate-compact.b2z"
# Save the in-memory table to a directory-backed .b2d store
climate.save(disk_path)
print(f"Saved to '{disk_path}'")
total_kb = sum(os.path.getsize(os.path.join(r, f)) for r, _, fs in os.walk(disk_path) for f in fs) / 1024
print(f"On-disk size: {total_kb:.1f} KB")
print(f"In-memory compressed: {climate.cbytes / 1024:.1f} KB")
Saved to '/var/folders/tb/7hwq2y354bb_68xwxjwjwwlr0000gn/T/blosc2_climate_pnnzna2z/climate'
On-disk size: 55.9 KB
In-memory compressed: 53.4 KB
Fast conversion between .b2d and .b2z¶
to_b2z() and to_b2d() use fast physical pack/unpack paths when possible: already-compressed leaves are copied as-is, without recompressing columns. This preserves the physical layout, including deleted rows and spare capacity.
Use compact=True when you want a logical compacted copy containing only visible/live rows. That path may rewrite columns and is slower.
[ ]:
# Fast-pack .b2d -> .b2z
ro = CTable.open(disk_path, mode="r")
ro.to_b2z(zip_path, overwrite=True)
ro.close()
print(f"Packed into '{zip_path}'")
# Fast-unpack .b2z -> .b2d
zipped = CTable.open(zip_path, mode="r")
zipped.to_b2d(unpacked_path, overwrite=True)
zipped.close()
print(f"Unpacked into '{unpacked_path}'")
# Logical compacted copy
ro = CTable.open(disk_path, mode="r")
ro.to_b2z(compact_zip_path, overwrite=True, compact=True)
ro.close()
print(f"Compacted copy: '{compact_zip_path}'")
[27]:
# Open read-only — fast, no data is copied until you access a column
ro = CTable.open(disk_path, mode="r")
print(f"Opened read-only: {len(ro):,} rows")
print(f"Cairo annual mean: {ro.where(ro.city == 'Cairo').temperature.mean():.1f} °C")
ro.close()
# Load fully into RAM (useful when you need repeated random access)
ram = CTable.load(disk_path)
print(f"Loaded into RAM : {len(ram):,} rows")
shutil.rmtree(tmpdir)
print("Temporary files removed.")
Opened read-only: 3,650 rows
Cairo annual mean: 21.9 °C
Loaded into RAM : 3,650 rows
Temporary files removed.
Part 8 — Arrow & CSV interop¶
CTable speaks Arrow and CSV, so it fits naturally into data pipelines.
[28]:
# CTable → Arrow
arrow_table = climate.select(["city", "day", "temperature"]).to_arrow()
print("Arrow table schema:", arrow_table.schema)
print("First 3 rows:", arrow_table.slice(0, 3).to_pydict())
Arrow table schema: city: string
day: int16
temperature: float
First 3 rows: {'city': ['Madrid', 'Madrid', 'Madrid'], 'day': [1, 2, 3], 'temperature': [2.909233808517456, 0.17393270134925842, 3.712681531906128]}
[29]:
import os
import tempfile
# CTable → CSV → CTable round-trip
tmp_csv = tempfile.mktemp(suffix=".csv")
climate.select(["city", "day", "temperature", "humidity"]).to_csv(tmp_csv)
print(f"CSV size: {os.path.getsize(tmp_csv) / 1024:.1f} KB")
@dataclass
class SlimReading:
city: str = blosc2.field(blosc2.string(max_length=16))
day: int = blosc2.field(blosc2.int16(ge=1, le=365), default=1)
temperature: float = blosc2.field(blosc2.float32(), default=0.0)
humidity: float = blosc2.field(blosc2.float32(), default=0.0)
t_from_csv = CTable.from_csv(tmp_csv, SlimReading)
print(f"Loaded from CSV: {len(t_from_csv):,} rows")
os.remove(tmp_csv)
CSV size: 111.5 KB
Loaded from CSV: 3,650 rows
Part 9 — Nullable columns¶
Real-world data is often incomplete. CTable handles missing values through a null sentinel approach: you declare a specific value (e.g. -1, "", or float("nan")) that represents “no data” for a column. The library treats it transparently in aggregates, sorting, unique(), value_counts(), and Arrow export.
This pattern is most useful for integer and string columns, which have no natural missing-value representation (unlike floats, which can use NaN).
[30]:
from dataclasses import dataclass
# Sensor log where some readings may be missing
@dataclass
class SensorLog:
sensor_id: int = blosc2.field(blosc2.int32(ge=0))
# -999 means "offline" — bypasses the ge/le constraint when stored
temperature: float = blosc2.field(blosc2.float64(ge=-50.0, le=60.0, null_value=-999.0), default=-999.0)
# "" means "location unknown"
location: str = blosc2.field(blosc2.string(max_length=16, null_value=""), default="")
log_data = [
(0, 22.3, "roof"),
(1, -999.0, "cellar"), # temperature: offline
(2, 18.7, ""), # location: unknown
(3, 31.5, "garage"),
(4, -999.0, ""), # both missing
(5, 15.1, "roof"),
]
log = blosc2.CTable(SensorLog, new_data=log_data)
print(log)
sensor_id temperature location
int32 float64 U16 (Unicode, max 16 chars)
──────────── ───────────────── ─────────────────────────────
0 22.3 roof
1 -999.0 cellar
2 18.7
3 31.5 garage
4 -999.0
5 15.1 roof
──────────── ───────────────── ─────────────────────────────
6 rows × 3 columns
9.1 Detecting nulls¶
[31]:
# is_null() → boolean array aligned to live rows
print("temperature is_null:", log["temperature"].is_null().tolist())
print("location is_null :", log["location"].is_null().tolist())
print()
print(f"Offline sensors (null temperature): {log.temperature.null_count()}")
print(f"Unknown locations : {log['location'].null_count()}")
# Use notnull() as a mask to select only valid readings
valid_temps = log["temperature"][:][log["temperature"].notnull()]
print(f"Valid temperature readings: {valid_temps}")
temperature is_null: [False, True, False, False, True, False]
location is_null : [False, False, True, False, True, False]
Offline sensors (null temperature): 2
Unknown locations : 2
Valid temperature readings: [22.3 18.7 31.5 15.1]
9.2 Null-aware aggregates¶
[32]:
# All aggregates automatically skip the null sentinel
temp = log["temperature"]
print(f"mean = {temp.mean():.2f} (3 valid readings only)")
print(f"min = {temp.min():.2f}")
print(f"max = {temp.max():.2f}")
print()
# unique() and value_counts() also exclude the sentinel
print("location unique:", log["location"].unique().tolist())
mean = 21.90 (3 valid readings only)
min = 15.10
max = 31.50
location unique: ['cellar', 'garage', 'roof']
9.3 Validation bypass¶
[33]:
# The sentinel bypasses ge/le constraints — you can store it freely
# even though -999.0 is below ge=-50.0
log.append((6, -999.0, "attic")) # succeeds: -999 is the null sentinel
print(f"Rows after append: {len(log)}")
# A genuine constraint violation still raises:
try:
log.append((7, -100.0, "lab")) # -100 is NOT the sentinel → rejected
except ValueError:
print("Rejected: temperature -100 violates ge=-50")
Rows after append: 7
Rejected: temperature -100 violates ge=-50
9.4 Sort: nulls always go last¶
[34]:
# Regardless of ascending / descending, null rows are placed at the end
s = log.sort_by("temperature")
print("Ascending (nulls last):")
print([round(v, 1) for v in s["temperature"][:].tolist()])
s_desc = log.sort_by("temperature", ascending=False)
print("Descending (nulls still last):")
print([round(v, 1) for v in s_desc["temperature"][:].tolist()])
Ascending (nulls last):
[15.1, 18.7, 22.3, 31.5, -999.0, -999.0, -999.0]
Descending (nulls still last):
[31.5, 22.3, 18.7, 15.1, -999.0, -999.0, -999.0]
9.5 Arrow export: sentinels become Arrow nulls¶
[35]:
try:
arrow = log.to_arrow()
tc = arrow.column("temperature")
lc = arrow.column("location")
print(f"Arrow temperature null_count: {tc.null_count}")
print(f"Arrow location null_count : {lc.null_count}")
print("Arrow temperature values:", tc.to_pylist())
except Exception as e:
print(f"(pyarrow not available: {e})")
Arrow temperature null_count: 3
Arrow location null_count : 2
Arrow temperature values: [22.3, None, 18.7, 31.5, None, 15.1, None]
9.6 CSV: empty cells become the null sentinel¶
[36]:
import os
import tempfile
csv_content = """sensor_id,temperature,location
10,25.1,lab
11,,office
12,18.3,
"""
with tempfile.NamedTemporaryFile(mode="w", suffix=".csv", delete=False) as f:
f.write(csv_content)
csv_path = f.name
log2 = blosc2.CTable.from_csv(csv_path, SensorLog)
print(log2)
print(f"temperature null_count: {log2.temperature.null_count()}")
print(f"location null_count : {log2['location'].null_count()}")
os.unlink(csv_path)
sensor_id temperature location
int32 float64 U16 (Unicode, max 16 chars)
──────────── ───────────────── ─────────────────────────────
10 25.1 lab
11 -999.0 office
12 18.3
──────────── ───────────────── ─────────────────────────────
3 rows × 3 columns
temperature null_count: 1
location null_count : 1
Summary¶
Here’s everything we covered:
Feature |
API |
|---|---|
Create |
|
Insert |
|
View |
|
Filter |
|
Project |
|
Sort |
|
Aggregates |
|
Stats |
|
Mutate |
|
Persist |
|
Interop |
|
Nullable |
|
CTable is designed for compressed analytical workloads — large tables that need to stay small in RAM while still being fast to query and easy to persist.