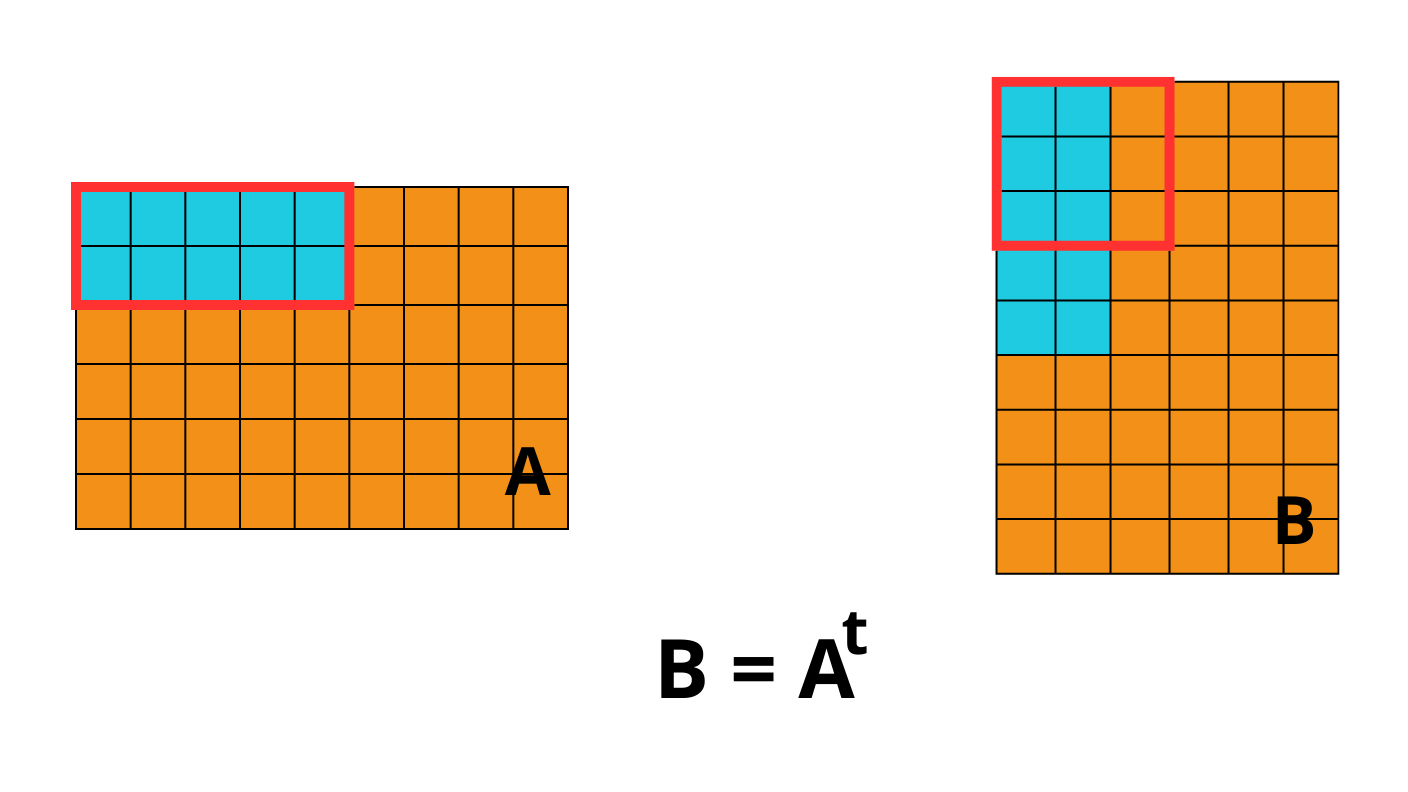
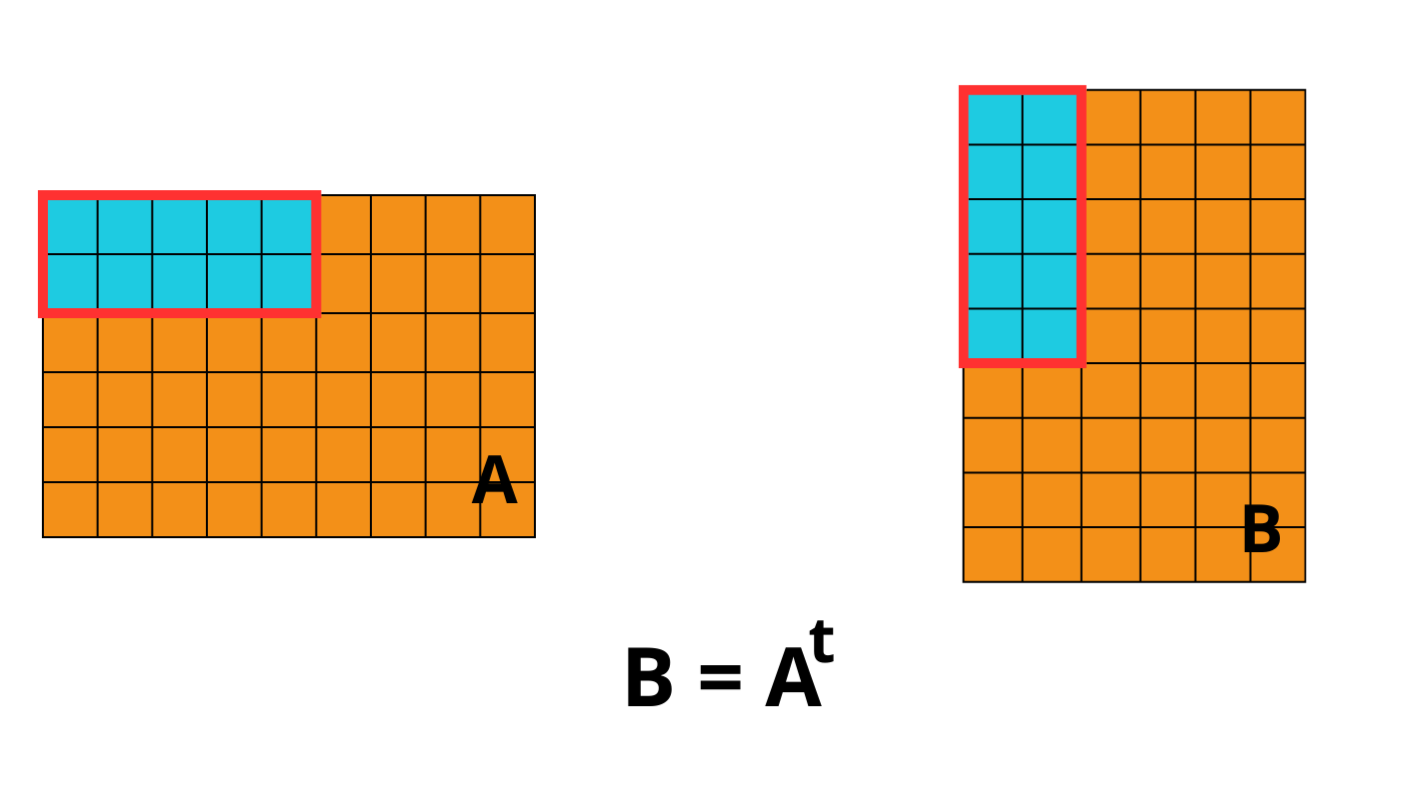
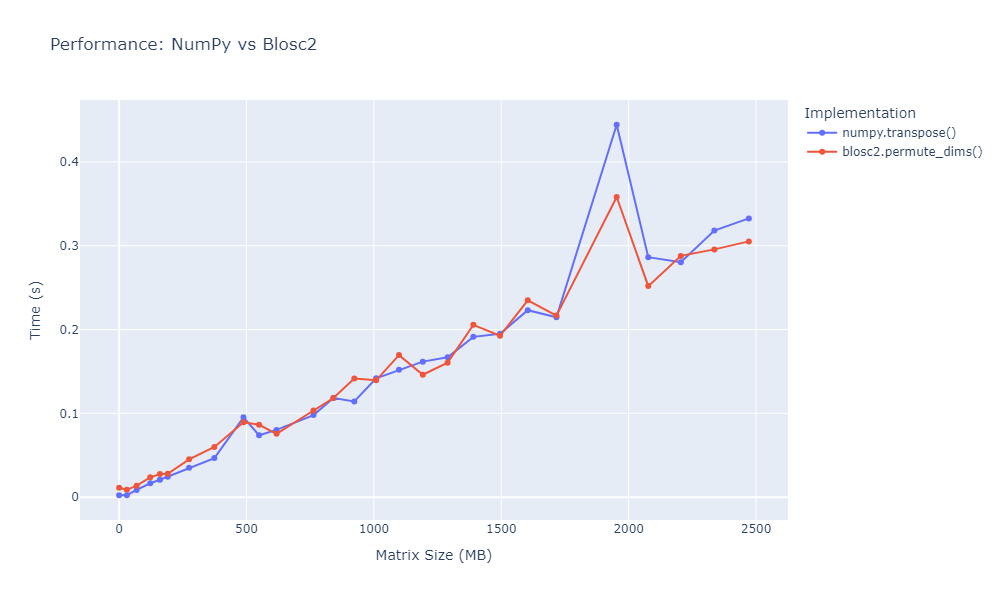
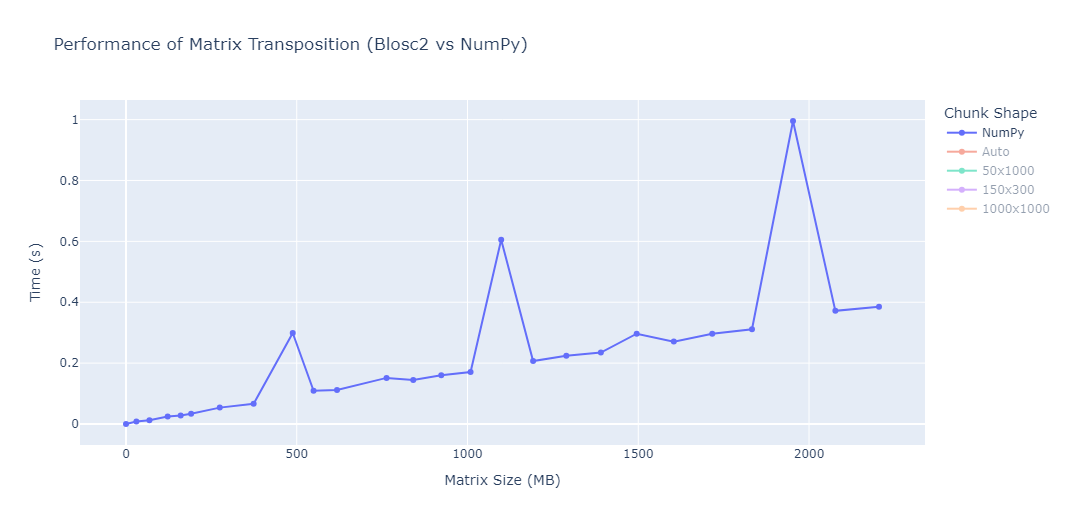
OpenZL Plugin for Blosc2
Blosc's philosophy of meta-compression is incredibly powerful - one is able to compose pipelines to optimally compress data (for speed or compression ratio), store information about the pipeline alognside the data in metadata, and then rely on a generic decompressor to read this and reverse the pipeline. The OpenZL team share our belief in the validity of this approach and have designed a graph-based formalisation with extensive support for all kinds of compression pipelines for all kinds of data.
However, Blosc2 is now much more than just a compression library - it offers comprehensive indexing support (including fancy indexing via the python-blosc2 interface) as well as an increasingly rapid compute engine (see this blog!). What if we could marry the incredibly comprehensive compression coverage of OpenZL with Blosc2's extended array manipulation functionality?
Foreseeing precisely this sort of challenge, prior Blosc2 developers implemented a dynamic plugin register functionality (loading the plugin in C-Blosc2, which can be called via Python-Blosc2). This means that with some unintrusive, relatively concise interface code, one can link Blosc2 and OpenZL at runtime (without substantially modifying either) and offer Blosc2 arrays compressed and decompressed with OpenZL.
The OpenZL plugin
The source code for the plugin can be found here. The minimal skeleton for the plugin layout follows
├── CMakeLists.txt
├── blosc2_openzl
│ └── __init__.py
├── pyproject.toml
├── requirements-build.txt
└── src
├── CMakeLists.txt
├── blosc2_openzl.c
└── blosc2_openzl.h
The blosc2_openzl.c must implement an encoder and decoder which are exported via an info struct:
#include "blosc2_openzl.h"
BLOSC2_OPENZL_EXPORT codec_info info = {
.encoder=(char *)"blosc2_openzl_encoder",
.decoder=(char *)"blosc2_openzl_decoder"
};
int blosc2_openzl_encoder(const uint8_t* src, uint8_t* dest,
int32_t size, uint8_t meta,
blosc2_cparams *cparams, uint8_t id) {
// code
}
int blosc2_openzl_decoder(const uint8_t *input, int32_t input_len, uint8_t *output,
int32_t output_len, uint8_t meta, blosc2_dparams *dparams,
const void *chunk) {
// code
}
The header blosc2_openzl.h then makes the info and encoder/decoder functions available to Blosc2:
#include "blosc2.h" #include "blosc2/codecs-registry.h" #include "openzl/openzl.h" BLOSC2_OPENZL_EXPORT int blosc2_openzl_encoder(...); BLOSC2_OPENZL_EXPORT int blosc2_openzl_decoder(...); // Declare the info struct as extern extern BLOSC2_OPENZL_EXPORT codec_info info;
PEP 427 and wheel structure
In order for the plugin to dynamically link to Blosc2, it has to be able to find the Blosc2 library at runtime. This has historically been quite finicky since different platforms and package managers may store Python packages (and the associated .so/.dylib/.dll library objects differently). Consequently, PEP 427 recommends distributing the Python wheels for packages which depend on compiled objects such as Python-Blosc2 in the following way
blosc2
├── __init__.py
├── lib
│ ├── libblosc2.so
│ ├── cmake
│ └── pkgconfig
└── include
└── blosc2.h
Finding the necessary libblosc2.so object from the top-level CMakeLists.txt file for the plugin is then as easy as:
# Find blosc2 package location using Python
execute_process(
COMMAND "${Python_EXECUTABLE}" -c "import blosc2, pathlib; print(pathlib.Path(blosc2.__file__).parent)"
OUTPUT_VARIABLE BLOSC2_PACKAGE_DIR
)
set(BLOSC2_INCLUDE_DIR "${BLOSC2_PACKAGE_DIR}/include")
set(BLOSC2_LIB_DIR "${BLOSC2_PACKAGE_DIR}/lib")
After building the plugin backend in src/CMakelists.txt one simply links the plugin to the backend (in this case openzl) and installs like so:
add_library(blosc2_openzl SHARED blosc2_openzl.c)
target_include_directories(blosc2_openzl PUBLIC ${BLOSC2_INCLUDE_DIR})
target_link_libraries(blosc2_openzl ${OPENZL_TARGET})
# Install
install(TARGETS blosc2_openzl
RUNTIME DESTINATION blosc2_openzl
LIBRARY DESTINATION blosc2_openzl
)
Note that it is not necessary to link blosc2_openzl and blosc2 in target_link_libraries as the former depends only on macros and structs defined in header files - and not functions. This makes the libblosc2_openzl.so object especially light and robust, as blosc2 is not registered as an explicit dependency. In fact on Linux, even if the blosc2_openzl.c were to include blosc2 functions, it is still not necessary to perform such linking!
Following PEP 427 allows one to add an additional safeguard to check if the plugin fails to find blosc2 by adding the RUNTIME_PATH property to the installed object
set_target_properties(blosc2_openzl PROPERTIES
INSTALL_RPATH "$ORIGIN/../blosc2/lib"
)
It also allows one to easily find the plugin .so object when calling via python - in the blosc2_openzl/__init__.py file one can find the library path as easily as os.path.abspath(Path(__file__).parent / libname) where libname is the desired .so/.dylib/.dll object (depending on platform). All these benefits have led us to update the wheel structure for python-blosc2 in the latest 4.0 release.
Using OpenZL from Python
Installing is then as simple as:
pip install blosc2_openzl
One can also download the project and use the cmake and cmake --build commands to compile C-level tests or examples. But let's get compressing with python straight away:
import blosc2
import numpy as np
import blosc2_openzl
from blosc2_openzl import OpenZLProfile as OZLP
prof = OZLP.OZLPROF_SH_BD_LZ4
# Define the compression parameters for Blosc2
cparams = {'codec': blosc2.Codec.OPENZL, 'codec_meta': prof.value}
# Create (uncompressed) array
np_array = np.arange(1000).reshape((10,100))
# Compression with the OpenZL codec
bl_array = blosc2.asarray(np_array, cparams=cparams)
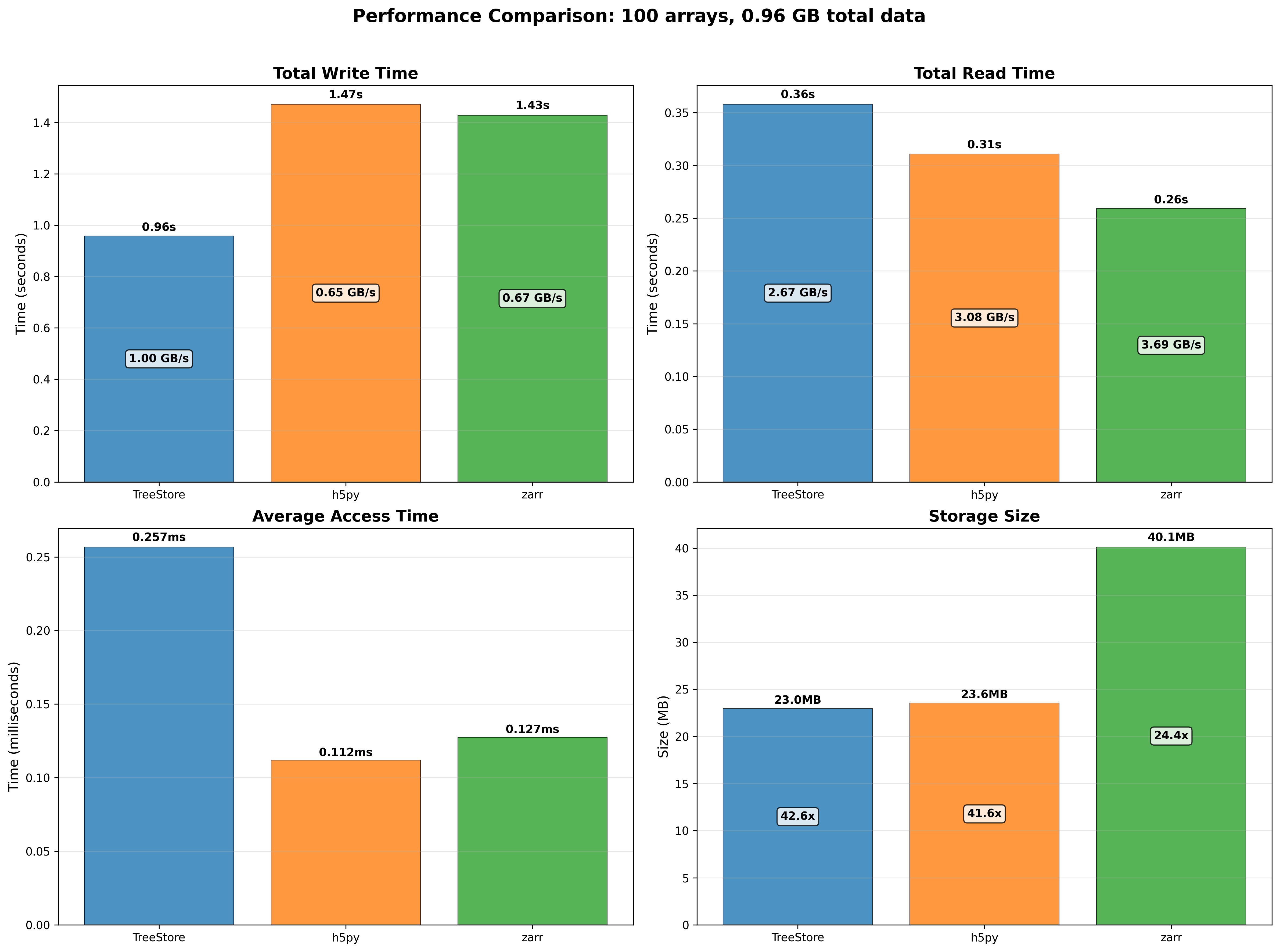
print(bl_array.cratio) # print compression ratio
## 25.078369905956112
The OpenZLProfile enum contains the available profile pipelines that have been implemented in the plugin, which use the codec_meta field (an 8-bit integer) to specify the desired transformation via codecs, filters and other nodes for the compression graph. Starting from the Least-Significant-Bit (LSB), setting the bits tells OpenZL how to build the graph:
CODEC | SHUFFLE | DELTA | SPLIT | CRC | x | x | x |
CODEC - If set, use LZ4. Else ZSTD.
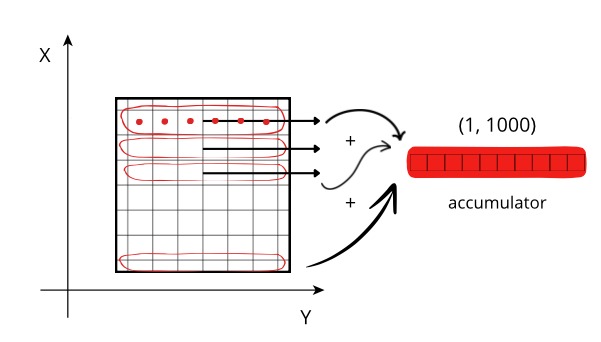
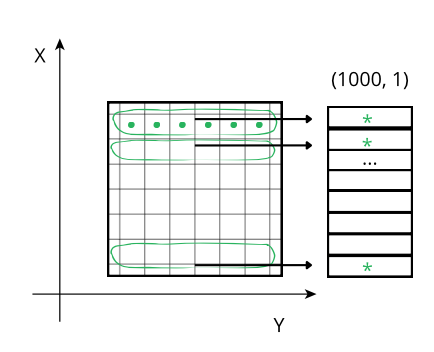
SHUFFLE - If set, use shuffle (outputs a stream for every byte of input data typesize)
DELTA - If set, apply a bytedelta (to all streams if necessary)
SPLIT - If set, do not recombine the byte streams
CRC - If set, store a checksum during compression and check it during decompression
The remaining bits may be used in the future.
In the future it would be great to further expand the OpenZL functionalities that we can offer via the plugin, such as bespoke transformers trained via machine learning techniques - see the OpenZL page for a flavour of what can be done with the (still evolving) library.
Conclusions
C-Blosc2's ability to support dynamically loaded plugins allows the library to grow in features without increasing the size and complexity of the library itself. For more information about user-defined plugins, refer to this blog entry. We have put this to work to offer linkage with the rather complex OpenZL library with a relatively rapid turnaround from design to prototype to full release in around a month. This is thanks to prior hard work by open source contributors from Blosc but naturally also OpenZL - many thanks to all!
If you find our work useful and valuable, we would be grateful if you could support us by making a donation. Your contribution will help us continue to develop and improve Blosc packages, making them more accessible and useful for everyone. Our team is committed to creating high-quality and efficient software, and your support will help us to achieve this goal.